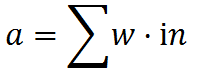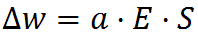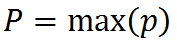Greetings!
The past 3 days I have spent on writing a fast GPU spatial and temporal pooler. It works! Unlike Numenta’s HTM, it both operates in the continuous domain and is much, much simpler. I don’t think I sacrificed any functionality by converting the discrete logic of the original into continuous formulas.
I sort of just made up formulas as I went, until it gave me the desired result. Not very rigorous, but it got the job done in the end regardless.
So, I would like to now describe the formulas that I came up with. Hopefully someone finds it useful besides myself. Continuous HTM is not well researched, if at all.
I will assume that you know how the original HTM works to some extent. For instance, what the spatial and temporal poolers are supposed to do.
I am not going to use proper indexing (e.g. wij as opposed to simply w) for formulas, I will rather describe in words how the operation is performed given the indexless formula. This is not supposed to be a paper, just a way of giving a rough idea of how the continuous spatial and temporal poolers work.
Continuous Spatial Pooler
The continuous spatial pooler consists of 2 2D buffers of floats, the activation buffer and the column output buffer.
The activation buffer receives input from another 2D buffer of floats, the input buffer. It has connections to a subregion of the input buffer for each element of the activation buffer. These subregions are positioned such that the normalized coordinates of the positions in both the input and activation buffers are the same.
One can then perform a weighted convolution of the input buffer. For each element in the activation buffer, compute:
Where a is the activation value, w is the weight of the connection, and in is the input corresponding to the connection. The activation value is stored in the activation buffer.
Once all activations have been computed, then comes the inhibition step. In this step, one reads the activation values surrounding the current element within a predefined radius, and finds the maximum. One then deactivates the current element according to how close it is to the maximum. So, something like this:
Where O is the output to be stored in the column output buffer, a is the activation of this element, m is the maximum activation across all elements in the inhibition radius, and c is a constant “intensity” factor.
This is the output of the spatial pooler. However, we are not done with it yet. We need to update the weights to learn unique sparse representations of the input. I used Oja’s rule for this:
From wikipedia:
Oja’s rule defines the change in presynaptic weights w given the output response  of a neuron to its inputs x to be
of a neuron to its inputs x to be
So, for each weight, we want to perform a hebbian-style learning rule. Oja’s rule is a version of the Hebbian rule that does not increase without bounds.
Temporal Pooler
Once we have the outputs of all of the elements of the column output buffer, it is time to compute the outputs of the cells within each column. Cells are represented by 2 3D buffers: The cell state buffer and the cell prediction buffer. The former contains the cell’s outputs, and the latter contains future predictions made by cells. We also have an additional column predictions buffer, which is 2D, and represents the predictions the cells gave to a column.
We first use a “soft” winner-takes-all scheme (so more like “winner takes most”) in order to give a higher output to cells which were previously predicted most accurately. This helps form a context for a column. The cells indicate which predictions activated this column into the predictive state, allowing future context-sensitive predictions to occur.
So, we first compute the prediction errors for each cell. This is simply the absolute value of the difference between the current column output O, and the cell predictions p from the previous frame:
While doing so, we keep track of the smallest error, which we will call x.
The output (state) of a cell S is then:
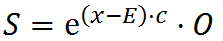
where c is again an “intensity” constant (doesn’t need to be equal to the previously mentioned c, it’s a different constant).
Once we have computed all cell activations, it is time to update the weights a cell has to all other cells in a radius. These are called lateral connections. They are updated according to their cell’s prediction error E as well as the state of the source cell from the previous time step:
Once we have updated the weights, it is time to make our prediction of the next frame’s input. This is a function of the summation of the weights multiplied by their respective cells’ states:
where w is in this case the cell lateral weights, S is the cell state to which the weight corresponds, and c is again an “intensity” constant, which can be different from the previously mentioned c‘s.
Finally, we create a column prediction from the cell predictions. This is simply the highest prediction in the column:
where P is the column’s prediction.
The Result
I would make a video, but I fear it would be too difficult to explain what it means. So instead, I will release a demo program along with source code within the next few days. Until then, you will have to take my word for it that it indeed works! I will edit this post to contain the link as soon as it is available.
Edit: The link: https://github.com/222464/ContinuousHTMGPU
How is this useful for outperforming Deepmind?
I have a plan on how to elegantly combine this continuous HTM with reinforcement learning, based on concepts discussed in my previous post. I will explain it in the next post, since this one has become quite long already.
See you next time!